20 Output
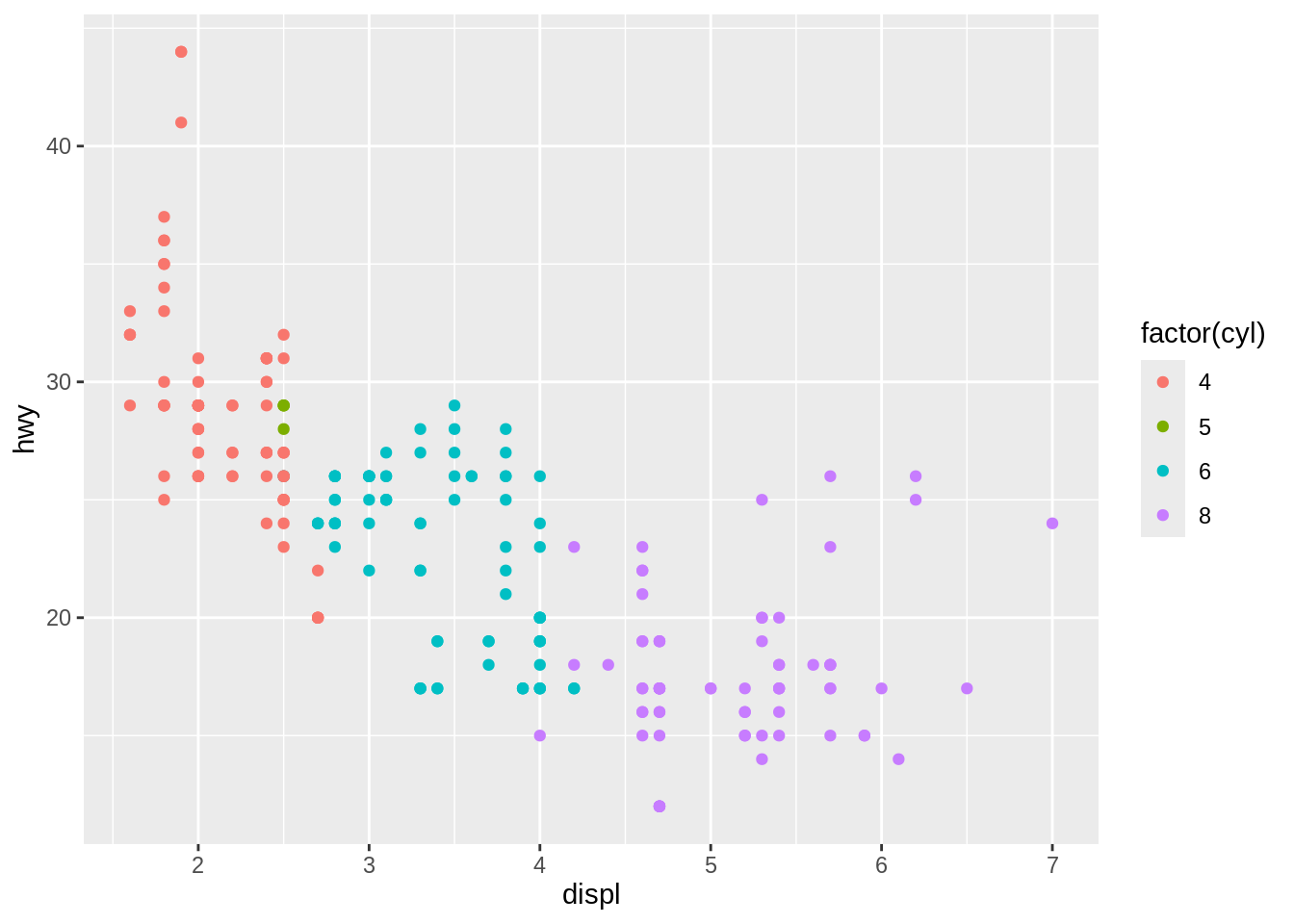
Most of the time you create a plot object and immediately render it, but you can also save a plot to a variable and manipulate it:
Once you have a plot object, there are a few things you can do with it:
- Render it on screen by calling its name or with
print(). This happens automatically when running interactively, but inside a loop or function, you’ll need toprint()it yourself.
- Save it to disk with
ggsave().
- Briefly describe its structure with
summary().
data: manufacturer, model, displ, year, cyl, trans, drv, cty, hwy, fl,
class [234x11]
mapping: x = ~displ, y = ~hwy, colour = ~factor(cyl)
faceting: <ggproto object: Class FacetNull, Facet, gg>
compute_layout: function
draw_back: function
draw_front: function
draw_labels: function
draw_panels: function
finish_data: function
init_scales: function
map_data: function
params: list
setup_data: function
setup_params: function
shrink: TRUE
train_scales: function
vars: function
super: <ggproto object: Class FacetNull, Facet, gg>
-----------------------------------
geom_point: na.rm = FALSE
stat_identity: na.rm = FALSE
position_identity - Save a cached copy of it to disk, with
saveRDS(). This saves a complete copy of the plot object, so you can easily re-create it withreadRDS().